AP Syllabus focus: ‘Polyprotic titration curves can reveal the number of acidic protons and identify major species and pKa values; quantitative speciation across the full curve is not assessed, but qualitative reasoning is.’
Polyprotic titrations extend the logic of monoprotic titrations by tracking multiple, stepwise neutralisations. The key skill is interpreting the titration curve qualitatively to infer how many protons are acidic and which species dominate in each region.
Polyprotic titrations: what you are looking for
A polyprotic acid donates more than one proton in separate steps, so it can be neutralised in stages by a strong base.
Polyprotic acid: an acid capable of donating two or more protons (H+) per formula unit in stepwise reactions.
In a titration of a polyprotic acid with a strong base, the curve often shows repeating patterns:
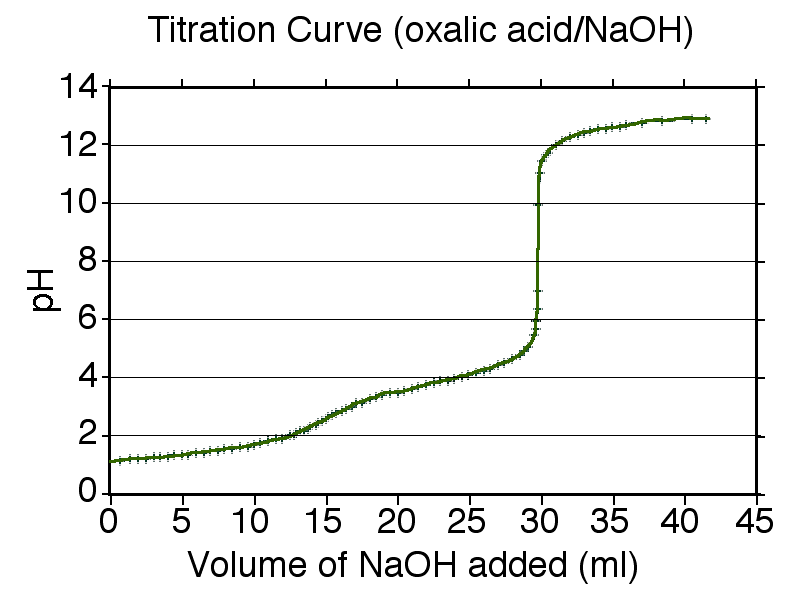
Diprotic acid titration curve (oxalic acid with NaOH) showing two distinct steep rises (two equivalence points) separated by a flatter buffer region. Visually, the repeated “flat → sharp rise → flat” pattern is the key evidence that more than one proton is titratable under the conditions of the titration. Source
Multiple buffer regions (relatively flat portions).
Multiple equivalence points (steep rises), potentially more than one.
Multiple half-equivalence points that align with different acidity steps.
Inferring the number of acidic protons (qualitative)
The number of titratable protons is revealed by how many distinct neutralisation stages are visible.
If the curve shows two clear steep rises (two equivalence points), the acid behaves as diprotic under those conditions.
If it shows three, it behaves as triprotic.
If later rises are not well separated, the acid may still be polyprotic, but the steps overlap so the curve looks “compressed.”
A quick stoichiometric idea connects equivalence points to how many protons have been neutralised, without needing full calculation detail.
= moles of OH− added at a chosen equivalence point, mol
= number of acidic protons neutralised up to that equivalence point (1 for first, 2 for second, etc.), unitless
= initial moles of polyprotic acid present, mol
This relationship supports qualitative reasoning such as “the second equivalence requires twice as many moles of base as the first.”
Identifying major species along the curve (qualitative)
Between key points on the curve, you should be able to name the predominant species (the form present in the largest amount).
Start (before much titrant is added)
Mostly the fully protonated form (for a diprotic acid, H2A dominates).
Small amounts of partially deprotonated forms may exist, but are not dominant.
After the first equivalence point (but before the second, for diprotic)
The partially deprotonated form (HA− for a diprotic acid) is typically the major species.
This intermediate can often act as both an acid and a base (amphiprotic behaviour), which helps explain why the pH is neither extremely acidic nor extremely basic in this region.
After the final equivalence point
Mostly the fully deprotonated conjugate base (A2− for diprotic, A3− for triprotic).
Any additional strong base added beyond the last equivalence makes the pH rise sharply due to excess OH−.
Connecting “steps” to pKa values (qualitative)
Each deprotonation step has its own pKa, so polyprotic titration curves can indicate multiple pKa values:
Each buffer region corresponds to a conjugate pair from one step (e.g., H2A/HA−, then HA−/A2−).
Each buffer region tends to centre around the acidity of that step, so the curve can be used to locate approximate pKa values.
What the AP Exam expects (and what it does not)
You are expected to:
Use the shape of a polyprotic titration curve to infer the number of acidic protons.
Identify the major species in broad regions (start, between equivalence points, after the final equivalence).
Link each step to a distinct pKa qualitatively.
You are not expected to:
Perform quantitative speciation across the full curve (tracking exact concentrations of every species at every point); the emphasis is on qualitative reasoning from the curve’s features.
FAQ
If successive dissociation steps are not well separated (similar $K_a$ values), the pH changes can overlap.
High concentrations and limited pH range on the graph can also compress later steps, making an equivalence point hard to distinguish.
Look for whether the curve shows consistent, repeated buffering behaviour and whether the steps appear stoichiometrically linked (second equivalence occurring after about double the base added compared with the first).
A mixture often produces less orderly regions and depends strongly on relative amounts of each acid.
They can be amphiprotic: able to donate a proton in one context and accept a proton in another.
This can make the pH between equivalence points less extreme than you might expect from a “pure base” or “pure acid” picture.
As protons are removed, the remaining species are progressively more negatively charged and typically less willing to lose another proton.
After the final equivalence, the fully deprotonated species can behave as a base by reacting with water, raising pH further.
Concentration of analyte and titrant
Strength (spacing) of the dissociation steps (differences between $pK_a$ values)
Quality of pH measurements and mixing
Volume scale: small equivalence volumes are harder to resolve
Practice Questions
Q1 (2 marks) A titration curve for an unknown acid shows two distinct steep rises and two equivalence points when titrated with NaOH(aq). What does this indicate about the acid, and what is the predominant acid-derived species at the first equivalence point?
Identifies the acid as diprotic / has two titratable protons. (1)
States the predominant species at the first equivalence is the singly deprotonated form, (or equivalent notation). (1)
Q2 (5 marks) A student titrates with strong base and observes three buffer regions and three equivalence points. Explain, using species names/formulae, what dominates (i) at the start, (ii) between the first and second equivalence points, and (iii) after the third equivalence point. Also state what the three buffer regions imply about values.
Start: predominantly . (1)
Between 1st and 2nd equivalence: predominantly (and recognises it arises after one neutralisation step). (1)
After 3rd equivalence: predominantly (before any excess OH− discussion). (1)
Notes buffer regions correspond to conjugate pairs for successive steps (e.g., , , ). (1)
Concludes there are three distinct values (one per step) indicated by the three buffer regions. (1)

