AP Syllabus focus:
‘Membrane proteins have hydrophilic and hydrophobic regions that determine how they embed in the bilayer and interact with cytosol or lipids.’
Membrane proteins are positioned within the phospholipid bilayer in predictable ways. Their orientation is dictated by chemistry, affects cell function, and must be established correctly during membrane assembly and trafficking.
Core idea: protein regions match membrane regions
A plasma membrane’s interior is hydrophobic, so proteins that reside in it must present hydrophobic surfaces to lipids while keeping hydrophilic regions exposed to water (cytosol or extracellular fluid).
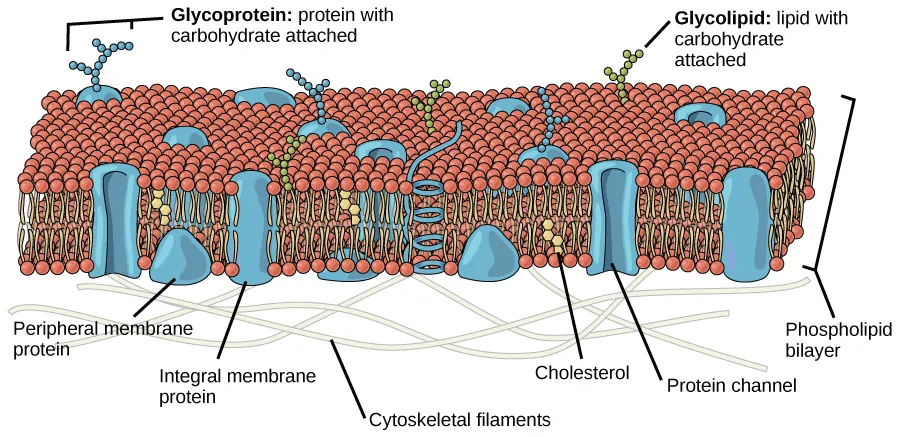
Fluid mosaic model diagram showing the phospholipid bilayer (hydrophobic core and hydrophilic surfaces) with embedded integral proteins and surface-associated peripheral proteins. The labeling makes it easy to connect protein placement to the chemistry of the membrane’s interior versus the aqueous cytosol/extracellular environment. Source
Amphipathic: having both hydrophobic and hydrophilic regions within the same molecule, allowing stable association with both membrane lipids and aqueous environments.
This amphipathic design explains why many membrane proteins have distinct “inside,” “membrane-spanning,” and “outside” portions.
Classes of membrane proteins by how they sit in the bilayer
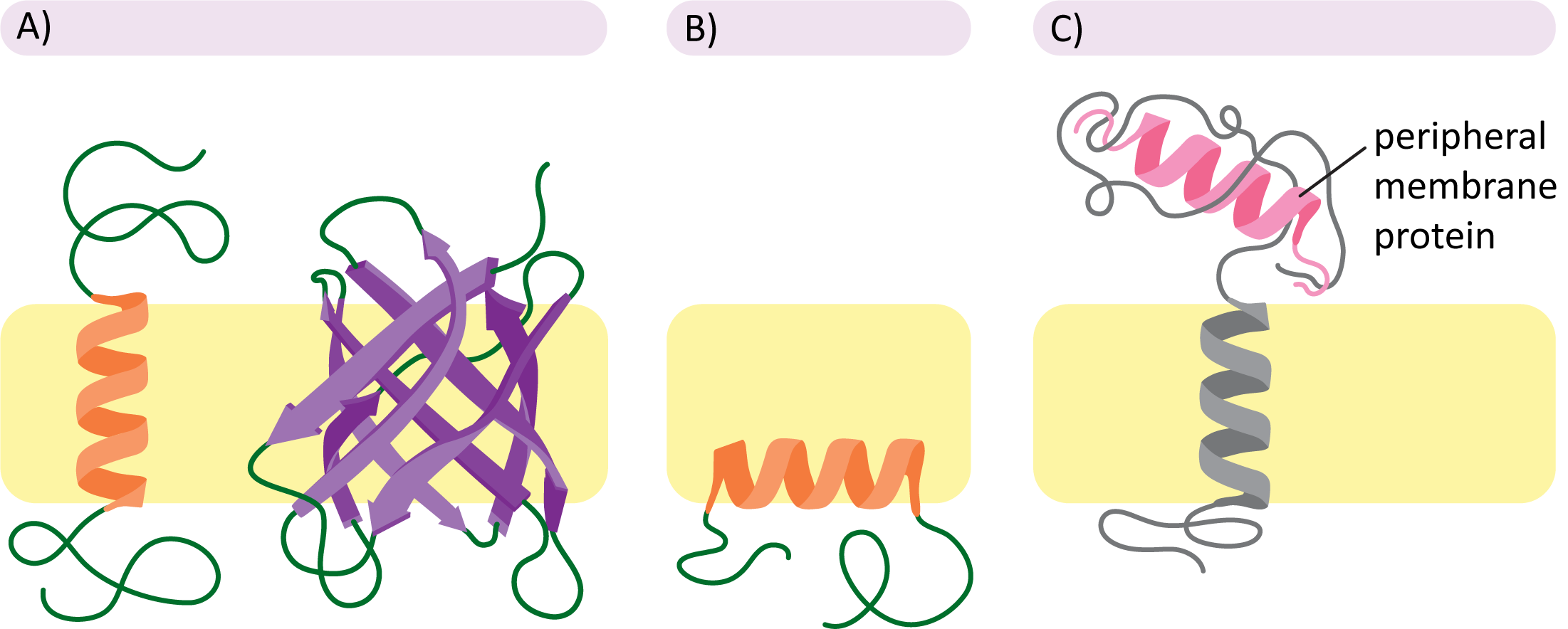
Examples of membrane protein classes: transmembrane integral proteins, monolayer-associated integral proteins, and peripheral proteins attached noncovalently at the membrane surface. Seeing these side-by-side emphasizes how hydrophobic segments stabilize integral proteins within lipid tails while peripheral proteins remain outside the hydrophobic core. Source
Integral (embedded) proteins
Integral membrane protein: a protein permanently associated with the membrane, with one or more hydrophobic regions embedded in the bilayer’s hydrophobic core.
Integral proteins often include transmembrane segments (commonly hydrophobic alpha helices) that anchor them in place.
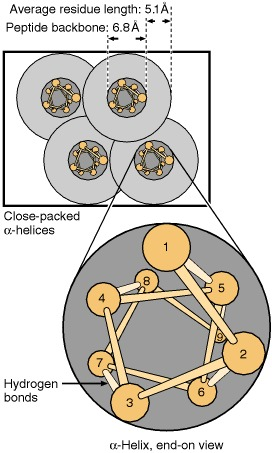
Figure showing that most transmembrane domains are α-helices, with amino-acid side chains projecting radially outward from the helix. This geometry helps explain why hydrophobic side chains can face the lipid core while the polypeptide backbone is stabilized by internal hydrogen bonding. Source
Single-pass proteins cross the membrane once.
Multi-pass proteins cross multiple times, creating distinct loops that face either the cytosol or the extracellular side.
Peripheral (surface-associated) proteins
Peripheral proteins do not penetrate the hydrophobic core. Instead, they associate with membranes by:
Binding to exposed regions of integral proteins
Associating with polar head groups of phospholipids
Forming ionic interactions with charged membrane surfaces
Because they remain on one side, peripheral proteins also have a meaningful sidedness (cytosolic-side vs extracellular-side).
What determines orientation (which side faces where)?
Hydrophobic “address labels” and stable insertion
Orientation is largely a consequence of which parts of the protein are hydrophobic enough to enter the bilayer and which parts are not.
Hydrophobic amino acids (e.g., leucine, isoleucine, valine, phenylalanine) favor the bilayer interior.
Polar/charged amino acids favor aqueous environments, so they tend to remain exposed to cytosol or extracellular fluid.
A protein’s final topology reflects the most energetically favorable arrangement: hydrophobic stretches buried in lipid tails, hydrophilic stretches exposed to water.
Directional interactions with cytosol and lipids
Once positioned, membrane proteins interact differently depending on which side they face:
Cytosolic-facing domains can bind cytoskeletal components or cytosolic enzymes, creating localized signaling or structural support.
Lipid-facing surfaces must be largely nonpolar to avoid destabilizing the bilayer.
Extracellular-facing domains often tolerate aqueous conditions and may participate in cell-cell recognition.
Why orientation matters for cell function
Correct orientation ensures that protein “active sites” and binding domains meet the correct partners.
Receptors must expose ligand-binding regions to the extracellular side while keeping signaling domains accessible to the cytosol.
Enzymes anchored to membranes act only where their substrates occur (cytosolic vs external), so sidedness controls pathway flow.
Cell recognition proteins present specific shapes outward; the same protein facing inward would be functionally useless for external identification.
Asymmetry in proteins contributes to overall membrane asymmetry: the two sides of a membrane are not interchangeable.
Maintaining orientation over time
Membrane protein orientation is stable because:
Hydrophobic segments are energetically “locked” into the bilayer core.
Lateral movement within a leaflet can occur, but flip-flopping of a protein across the bilayer is extremely unlikely without disassembly.
Cellular processes that add or remove membrane components must preserve which domains face the cytosol versus the non-cytosolic side.
FAQ
Methods include selective protease digestion of intact cells, antibody binding to non-permeabilised membranes, and chemical labelling that cannot cross the bilayer.
It graphs amino-acid hydrophobicity across a sequence. Long hydrophobic peaks suggest transmembrane segments, helping predict how many times a protein spans the bilayer.
Carbohydrate additions can stabilise folding, protect from proteases, and mediate recognition. Glycosylation patterns also help distinguish the non-cytosolic surface.
Not under normal conditions. Inversion would require moving hydrophilic regions through the hydrophobic core, which is energetically unfavourable without unfolding or specialised machinery.
Lipid rafts are microdomains enriched in certain lipids that can preferentially recruit specific proteins, influencing local orientation stability and clustering for signalling.
Practice Questions
Explain how hydrophobic and hydrophilic regions of a membrane protein determine its position in the phospholipid bilayer. (2 marks)
Hydrophobic amino acid regions embed within the hydrophobic core of the bilayer (1).
Hydrophilic/charged regions remain exposed to aqueous cytosol and/or extracellular fluid (1).
A student proposes that a receptor protein could function normally even if inserted into the plasma membrane with its domains reversed (extracellular domain facing cytosol). Evaluate this claim using principles of membrane protein orientation. (5 marks)
Membrane proteins have specific hydrophobic segments that must align with lipid tails to be stable (1).
Receptors require ligand-binding sites to be exposed on the extracellular side to bind external signals (1).
Cytosolic domains are needed to interact with intracellular signalling proteins/second messengers (1).
Reversing orientation would prevent correct interactions with either ligand or intracellular partners (1).
Orientation is energetically constrained and maintained; spontaneous inversion is highly unlikely (1).

