AP Syllabus focus: ‘In large biomolecules, noncovalent interactions may occur between different molecules or between different regions of the same large biomolecule.’
Large biomolecules rely on many weak attractions to create stable, functional shapes and assemblies. Understanding where these noncovalent interactions occur helps connect molecular-scale forces to properties like folding, binding, and material strength.
What “noncovalent” means in large biomolecules
Noncovalent interaction: An attractive (or repulsive) interaction that does not involve sharing electrons in a covalent bond; typically weaker than covalent bonds but significant when many act together.
Noncovalent interactions are individually small compared with covalent bonds, but biomolecules have many interaction sites, so the net effect can be large and highly structure-directing.
Common noncovalent interactions used by biomolecules
London dispersion forces (LDF): Present between all atoms; become especially important across large surface areas (many contact points).
Dipole–dipole attractions: Occur when polar groups align so partial charges attract.
Hydrogen bonding: Directional attractions involving H bonded to electronegative atoms and lone pairs on electronegative atoms; often crucial for shape recognition.
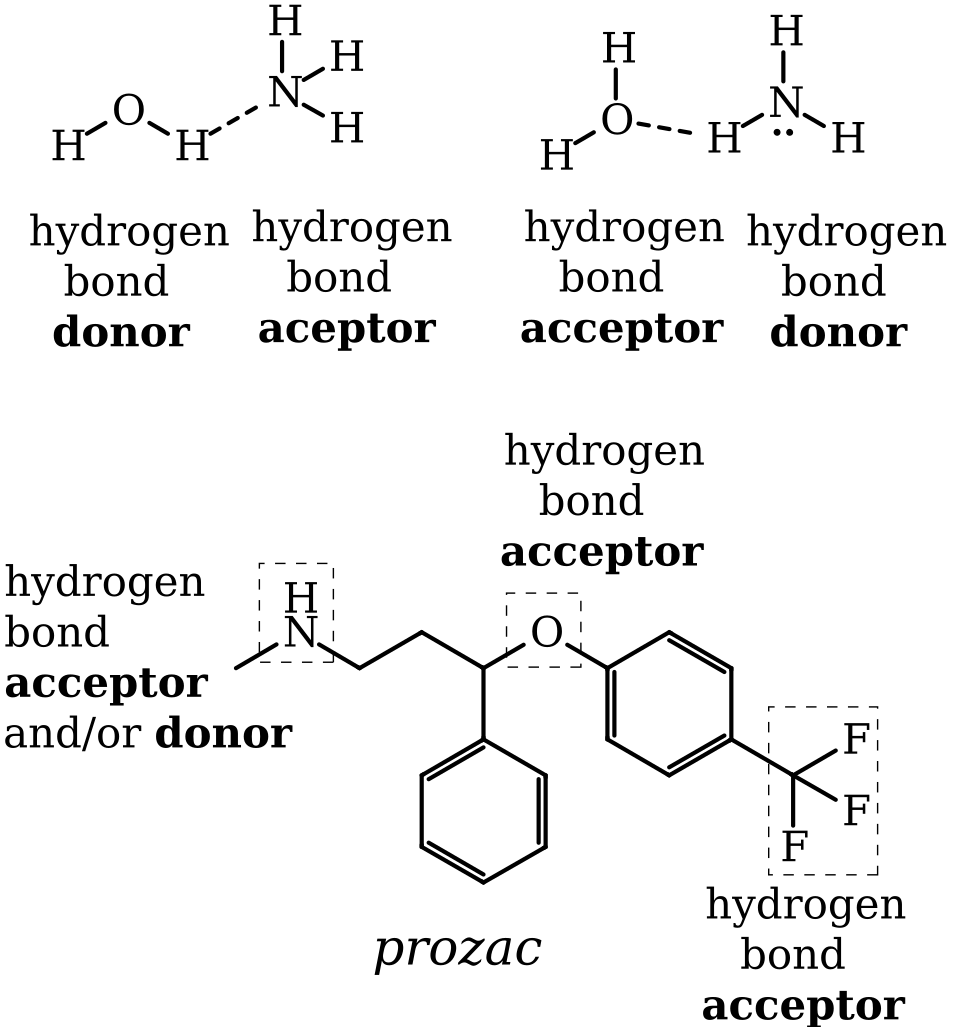
Diagram identifying hydrogen-bond donors and acceptors in common functional groups and small molecules. This helps you determine whether two nearby biomolecular sites can form a hydrogen bond (requires a donor X–H and an acceptor lone pair, arranged with the right geometry). Source
Electrostatic attractions (ionic “salt bridges” in biomolecules): Attractions between full charges on ionised side chains or groups.
Repulsions also matter: Like-charge repulsion or steric crowding can prevent certain shapes and favour others.
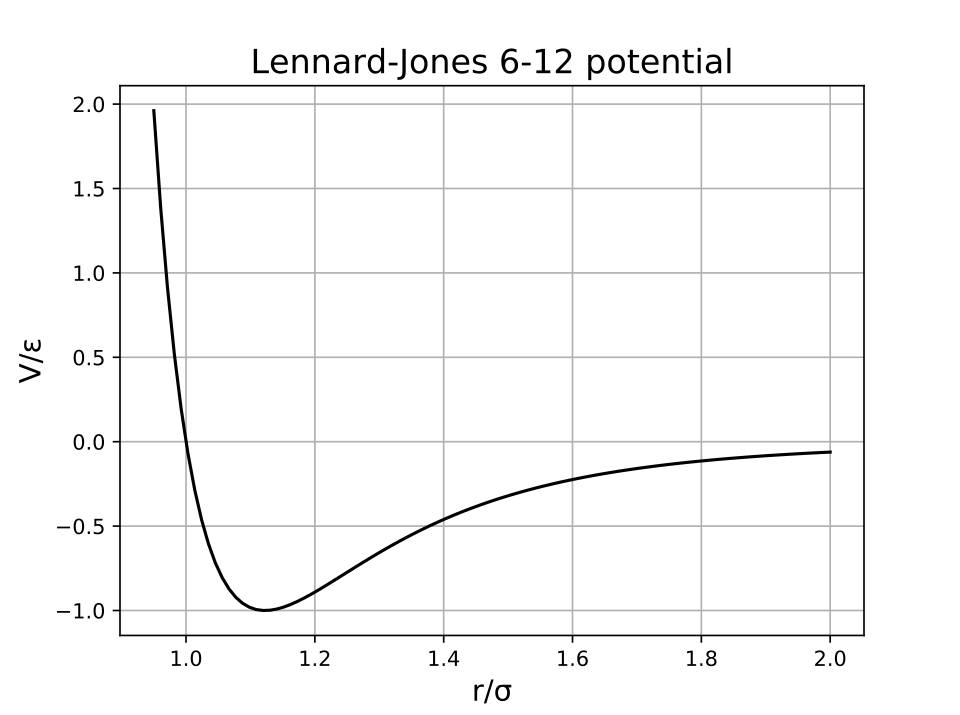
Lennard–Jones potential energy curve showing an attractive well at intermediate separation (dominated by dispersion, proportional to in the model) and a steep repulsive wall at short distances (electron-cloud overlap/Pauli repulsion). The minimum corresponds to the most energetically favorable contact distance, helping explain why tightly packed regions in folded biomolecules can be stable but cannot interpenetrate. Source
Noncovalent interactions within one large biomolecule (intramolecular)
The syllabus emphasises that noncovalent interactions can occur between different regions of the same large biomolecule. These long-range contacts stabilise specific 3D conformations.
Proteins: folding and internal organisation
Proteins have many polar and nonpolar regions; folding brings distant segments into close proximity.
Hydrogen bonds can stabilise repeated local patterns (for example, backbone interactions) and also help lock distant segments together.
LDF contribute wherever parts of the chain pack closely (many atoms in contact), making tight packing energetically favourable.
Electrostatic attractions between oppositely charged side chains can stabilise particular folds, while repulsions can destabilise them.
Nucleic acids: shape and recognition surfaces
Within DNA or RNA, hydrogen bonding and dipole interactions help maintain consistent geometries and create grooves with specific patterns of partial charge.
Stacking interactions between adjacent bases involve substantial LDF across large, flat surfaces, helping stabilise ordered structures.
Noncovalent interactions between biomolecules (intermolecular)
The syllabus also highlights noncovalent interactions between different molecules. Many biological functions are based on selective binding that is strong overall but reversible.
Molecular recognition (binding) is “many weak, well-placed forces”
Binding sites are shaped so that multiple attractions occur at once: hydrogen bonds, dipole alignment, and extensive LDF contact.
Specificity often comes from requiring the right spatial arrangement to satisfy several interactions simultaneously.
Reversibility is enabled because individual interactions are weak; dissociation becomes possible when enough contacts are disrupted.
Larger assemblies and materials
Biomolecular complexes (multi-protein assemblies, protein–nucleic acid complexes) are stabilised by networks of noncovalent interactions across interfaces.
When large surfaces meet, LDF can become a dominant contribution due to the sheer number of atom–atom contacts.
Noncovalent crosslinking between polymer-like biomolecules can influence macroscopic properties such as flexibility, viscosity, and toughness.
Interpreting biomolecule behaviour using intermolecular-force ideas
If conditions reduce attractive interactions (or increase repulsions), shapes and assemblies can change.
Because many interactions act together, a small change at multiple sites can significantly alter overall stability and function.
A useful particulate-level mindset: identify charged regions, polar groups capable of alignment, and broad contact surfaces where LDF can accumulate.
FAQ
By changing folding, a distant substitution can shift the positions of many groups that form noncovalent contacts.
Small geometric changes can weaken multiple interactions at once, lowering overall affinity.
Interfaces often involve large contact areas with many atoms close together.
Even though each dispersion interaction is weak, the summed effect across the interface can be substantial.
Water can compete for polar sites, weakening some direct intermolecular contacts.
It can also promote association of nonpolar surfaces by favouring reduced exposed nonpolar area.
Affinity describes how strongly two species bind overall.
Specificity describes how selectively a binding site prefers one partner over others, often due to shape and interaction-pattern matching.
Because the forces are noncovalent, components can dissociate and reassociate.
If the most stable arrangement maximises favourable contacts, the system can return to that state when conditions allow.
Practice Questions
Q1 (2 marks) A large biomolecule folds so that two distant nonpolar regions become tightly packed together. Identify the main type of noncovalent interaction stabilising this contact and justify your choice.
Identifies London dispersion forces / dispersion interactions (1)
Justifies with idea of many close atom–atom contacts and increased net attraction over a large surface area (1)
Q2 (5 marks) A protein binds to a short DNA segment. Describe how noncovalent interactions can (i) stabilise the protein’s folded shape and (ii) stabilise the protein–DNA complex, and explain why the overall binding can be strong yet reversible.
(i) Mentions intramolecular noncovalent interactions between different regions of the protein stabilising folding (1)
Correctly names at least one relevant interaction for folding (e.g. hydrogen bonds, electrostatic attractions, LDF) (1)
(ii) Mentions intermolecular noncovalent interactions between protein and DNA stabilising the complex (1)
Correctly links specificity to multiple well-aligned interactions/contact points (1)
Explains strong yet reversible as many individually weak interactions whose net is large but can be disrupted without breaking covalent bonds (1)

