AP Syllabus focus: ‘VSEPR predicts molecular shape by treating electron pairs as repelling through Coulombic interactions; use this idea to arrange electron domains around a central atom.’
VSEPR theory provides a fast, reliable way to predict 3D molecular shapes from a Lewis structure by arranging electron density to minimise repulsions and thus lower potential energy.
Core idea of VSEPR
Valence Shell Electron Pair Repulsion (VSEPR) treats regions of electron density around a central atom as repelling each other due to Coulombic interactions (like charges repel). The most stable arrangement maximises separation between these regions.
Electron domain: a region of electron density around a central atom (a single bond, multiple bond, or lone pair) that acts as one repelling unit in VSEPR.
Electron domains determine the overall electron-domain geometry; then you adjust to get the molecular shape (positions of atoms only).
Molecular shape (molecular geometry): the 3D arrangement of bonded atoms around a central atom, ignoring lone pairs.
How to apply VSEPR (process)
Step-by-step method
Draw a correct Lewis structure (central atom identified, valence electrons placed).
Count electron domains around the central atom:
Each single bond = 1 domain
Each multiple bond = 1 domain (electron density is in one direction)
Each lone pair = 1 domain
Arrange domains to maximise separation (minimise repulsion).
Name electron-domain geometry from total domains.
Convert to molecular shape by “hiding” lone pairs while keeping the same domain arrangement.
Relative repulsions (why angles change)
Lone pairs occupy more space than bonding pairs, so repulsions follow:
lone pair–lone pair > lone pair–bonding pair > bonding pair–bonding pair
This typically compresses bond angles compared with ideal values for a given electron-domain geometry.
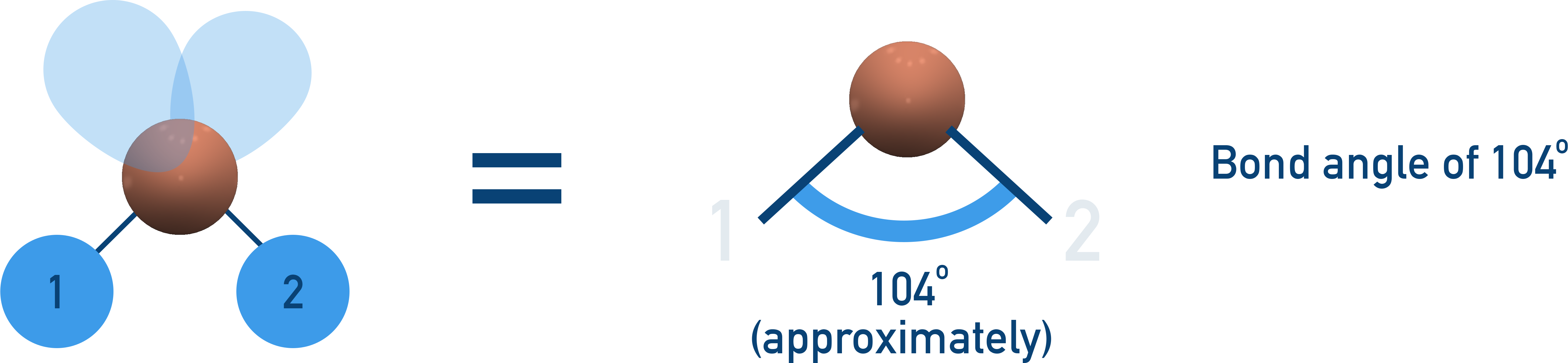
Bent molecular shape and angle compression: the diagram shows a bent molecule with a bond angle near 104°, illustrating how lone pairs (higher electron density) push bonding pairs closer together. This is why real molecules like H2O deviate from the ideal tetrahedral angle of 109.5°. Source
Common electron-domain geometries and resulting shapes
2 electron domains
Electron-domain geometry: linear
Molecular shape: linear (no lone-pair alternative for 2 domains)
Domains point opposite to maximise separation (180°).
3 electron domains
Electron-domain geometry: trigonal planar
Molecular shapes:
0 lone pairs: trigonal planar
1 lone pair: bent (lone pair increases repulsion, angle < 120°)
4 electron domains
Electron-domain geometry: tetrahedral
Molecular shapes:
0 lone pairs: tetrahedral
1 lone pair: trigonal pyramidal
2 lone pairs: bent (angles progressively smaller than 109.5°)
5 electron domains (trigonal bipyramidal arrangement)
Domains occupy equatorial (3 positions, 120° apart) and axial (2 positions, 180° apart, 90° to equatorial).
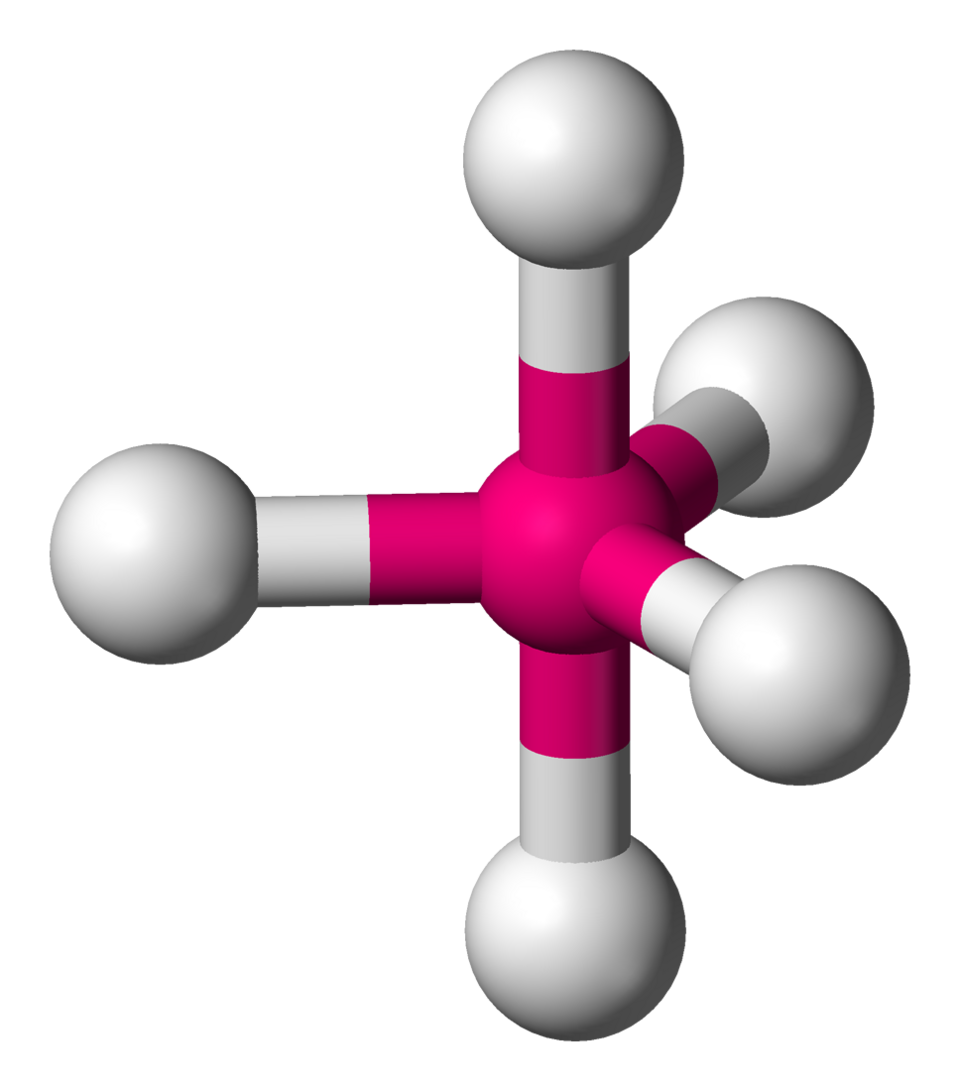
Trigonal bipyramidal electron-domain geometry (AX5E0): three domains form an equatorial plane (120° apart) and two occupy axial positions (180° apart). This framework explains why lone pairs preferentially take equatorial sites, reducing the number of 90° interactions. Source
Lone pairs preferentially occupy equatorial positions (fewer 90° interactions).
Molecular shapes depend on how many lone pairs replace bonded atoms in this framework (e.g., seesaw, T-shaped, linear).
6 electron domains (octahedral arrangement)
Electron-domain geometry: octahedral (90° separations)
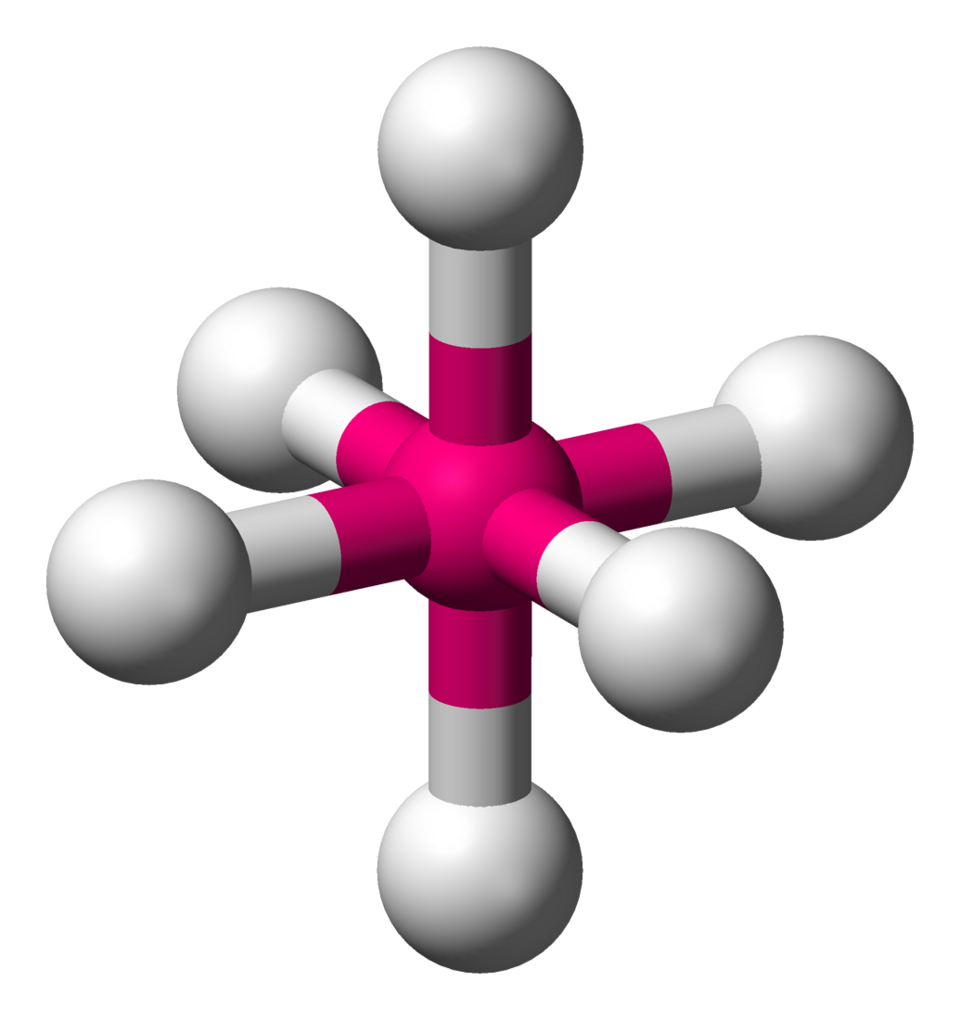
Octahedral electron-domain geometry (AX6E0): six bonding domains are arranged to maximize separation, giving 90° angles between adjacent positions and 180° between opposite positions. This is the idealized geometry used in VSEPR before considering any lone-pair substitutions (e.g., square pyramidal or square planar). Source
Lone pairs take positions to minimise 90° interactions:
1 lone pair: square pyramidal
2 lone pairs: square planar (lone pairs opposite)
What VSEPR does and does not assume
VSEPR is a model: it predicts shapes by repulsion minimisation, not by explicitly calculating energies.
It is most dependable for main-group molecules/ions where electron domains are well localised around one central atom.
FAQ
A multiple bond is counted as one electron domain because its electron density occupies one general direction from the central atom.
However, its higher electron density can increase repulsion slightly, sometimes causing small angle distortions.
Electron-domain geometry counts all domains (bonding and lone pairs) to set the 3D framework.
Molecular shape describes only where atoms are, so lone pairs change the name even though the domain framework is unchanged.
Different domains repel unequally (lone pairs spread out more), and surrounding atoms/groups can have different sizes.
These effects slightly adjust angles away from idealised symmetry-based values.
Placing the two lone pairs opposite (180° apart) minimises lone pair–lone pair repulsion.
The remaining four bonds then lie in one plane, giving a square planar arrangement of atoms.
Yes, apply VSEPR locally: treat each central atom separately using its own electron-domain count.
Overall molecular shape is then built from the connected local geometries.
Practice Questions
(1–3 marks) Phosgene, , has carbon as the central atom. Using VSEPR, state the electron-domain geometry around carbon and the molecular shape.
Counts 3 electron domains around C (C=O counts as 1, plus two C–Cl) (1)
Electron-domain geometry: trigonal planar (1)
Molecular shape: trigonal planar (1)
(4–6 marks) For a central atom with 5 electron domains, explain using VSEPR why lone pairs preferentially occupy equatorial positions rather than axial positions, and state one consequence for predicted molecular shape when one lone pair is present.
Identifies trigonal bipyramidal electron-domain geometry with axial and equatorial positions (1)
Links domain arrangement to minimising repulsions via Coulombic interactions/maximising separation (1)
States equatorial has fewer 90° interactions than axial (equatorial: two 90°; axial: three 90°) (2)
Concludes lone pair goes equatorial because lone pairs repel more strongly than bonding pairs (1)
Consequence: molecular shape becomes seesaw (or equivalent correct shape for one lone pair in trigonal bipyramidal) (1)

