AP Syllabus focus:
‘Ribosomes are non-membrane structures in all cells that synthesize proteins from mRNA, reflecting common ancestry and supporting cellular protein production.’
Ribosomes connect genetic information to cell function by translating mRNA into polypeptides. Their conserved structure across life highlights shared ancestry, and their placement in cells supports efficient protein production for diverse cellular needs.
Ribosomes as universal, non-membrane structures
Ribosomes are non-membrane complexes found in all cells (prokaryotic and eukaryotic). They are built from ribosomal RNA (rRNA) and proteins, emphasizing that RNA is central to gene expression and catalytic function.
Ribosome: A non-membrane rRNA–protein complex that translates mRNA codons into a polypeptide using tRNAs.
Because ribosomes are present in every living cell and their core features are strongly conserved, they support the AP idea of common ancestry: fundamental molecular machinery was inherited and modified over evolutionary time rather than independently invented in each lineage.
Ribosome structure and functional sites
Ribosomes have two subunits (large and small) that assemble on an mRNA during translation.
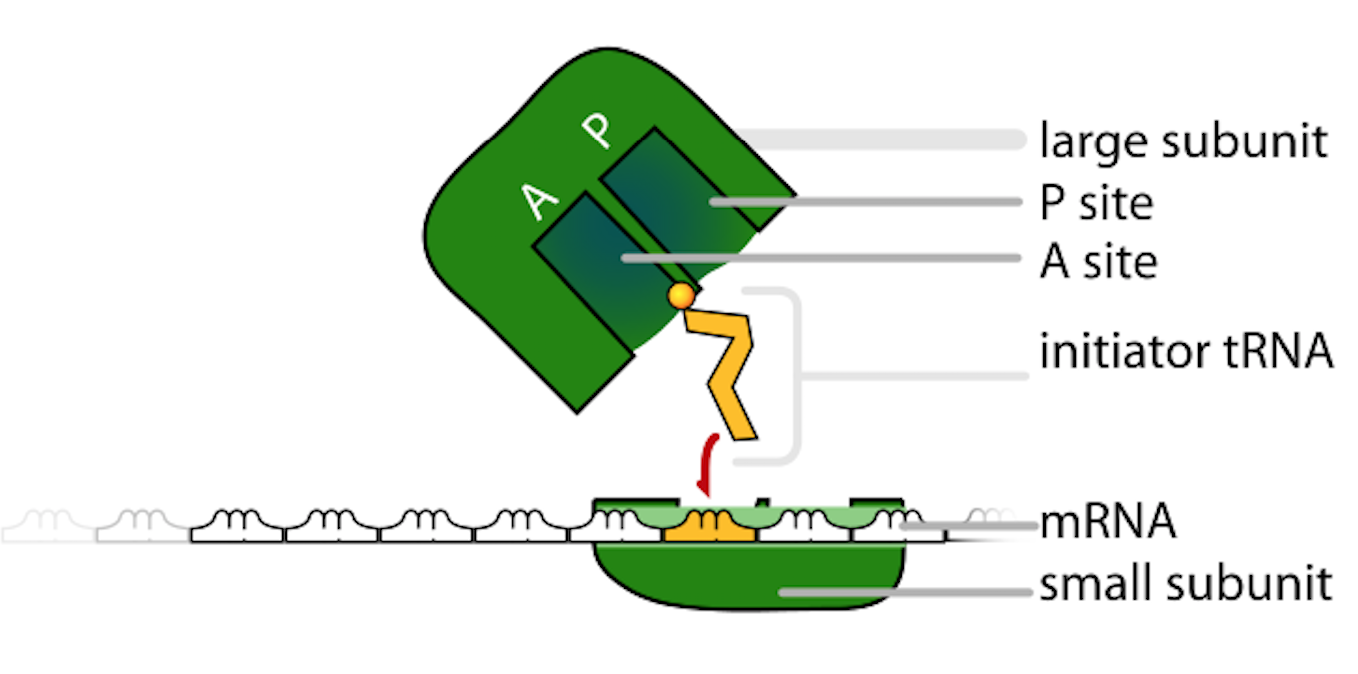
This diagram breaks down the ribosome into its large and small subunits and highlights the functional binding pockets used during translation. The labeled binding sites help connect ribosome structure to its role in positioning tRNAs and interpreting codons to build a polypeptide. Source
Key functional regions help enforce accurate protein synthesis:
mRNA-binding region: positions the message for reading
tRNA-binding sites (commonly A, P, and E sites): coordinate entry, peptide-bond formation, and exit of tRNAs
Catalytic center: rRNA participates directly in peptide-bond formation, making the ribosome a ribozyme in function
Ribosomes work as a coordinated machine: structure creates specificity (codon reading) while catalytic activity enables rapid polymer formation.
Protein synthesis from mRNA (translation)
Protein synthesis at ribosomes is the process of translation, in which the nucleotide sequence of mRNA is decoded into the amino acid sequence of a protein. The information flow depends on the genetic code, read as codons (triplets).
Codon: A three-nucleotide sequence in mRNA that specifies an amino acid or a stop signal during translation.
Translation depends on tRNA molecules that “interpret” codons by pairing an anticodon with the mRNA codon while carrying a specific amino acid.
Core requirements for translation
To synthesize proteins from mRNA, cells need:
An mRNA template with codons arranged 5’ to 3’
Ribosomal subunits and associated factors
tRNAs charged with amino acids (aminoacyl-tRNAs)
Amino acids as monomers for the polypeptide
Energy (commonly in the form of GTP/ATP equivalents) to drive initiation, elongation, and termination steps
Stages of translation at the ribosome
Translation is often described in three stages. The details may differ across organisms, but the logic is conserved.
Initiation: assembling the translation complex
The small ribosomal subunit binds the mRNA and identifies the start codon (usually AUG)
An initiator tRNA carrying methionine pairs with the start codon
The large subunit joins, establishing a functional ribosome with aligned tRNA-binding sites
Initiation sets the reading frame, ensuring codons are grouped correctly; a one-base shift changes every downstream codon and can drastically alter protein sequence.
Elongation: building the polypeptide chain
Elongation repeats a cycle that adds one amino acid per codon:
A charged tRNA enters the A site and base-pairs with the next codon
A peptide bond forms, transferring the growing chain to the amino acid in the A site
The ribosome translocates one codon forward, shifting tRNAs (A → P, P → E), and the empty tRNA exits
This stepwise movement allows the ribosome to synthesize proteins efficiently and accurately, producing a linear polypeptide that will later fold into a functional protein.
Termination: releasing the completed protein
When a stop codon (UAA, UAG, or UGA) enters the A site, no tRNA matches it
A release factor binds, promoting release of the polypeptide
Ribosomal subunits dissociate and can be reused
Termination ensures proteins have defined endpoints and prevents uncontrolled translation into noncoding regions.
Ribosomes and cellular protein production
Ribosomes support cellular function by generating proteins for structure, catalysis, transport, and regulation. Cells can adjust protein output by regulating:
mRNA availability (which messages are present)
Ribosome recruitment (how many ribosomes translate a given mRNA)
Translation efficiency (initiation frequency, elongation speed, and accuracy)
Free vs. membrane-associated ribosomes (functional significance)
Ribosomes can be found in different cellular locations, which supports targeted protein production:
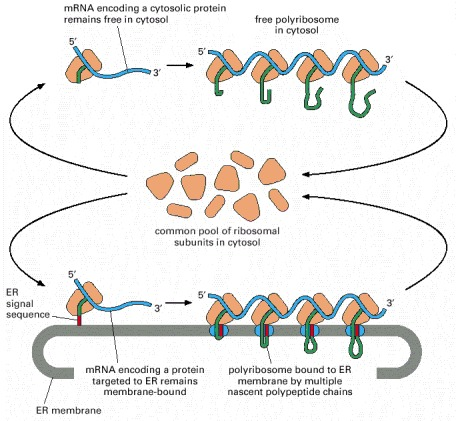
This textbook figure contrasts free ribosomes in the cytosol with ribosomes docked on the rough ER during synthesis of secreted or membrane proteins. It emphasizes that the ribosomes themselves come from a common pool; what changes is the presence of targeting information (e.g., an ER signal sequence) that routes translation to the ER. Source
Free ribosomes (in the cytosol) typically produce proteins that function in the cytosol
Ribosomes associated with membranes (e.g., attached to cytosolic faces of internal membranes) produce proteins that are directed to specific cellular destinations
The key AP idea is that ribosomes themselves are structurally similar; it is the destination of the protein and targeting information that differs, not a fundamentally different ribosome type.
Polyribosomes increase output
Multiple ribosomes can translate the same mRNA simultaneously, forming a polyribosome (polysome).
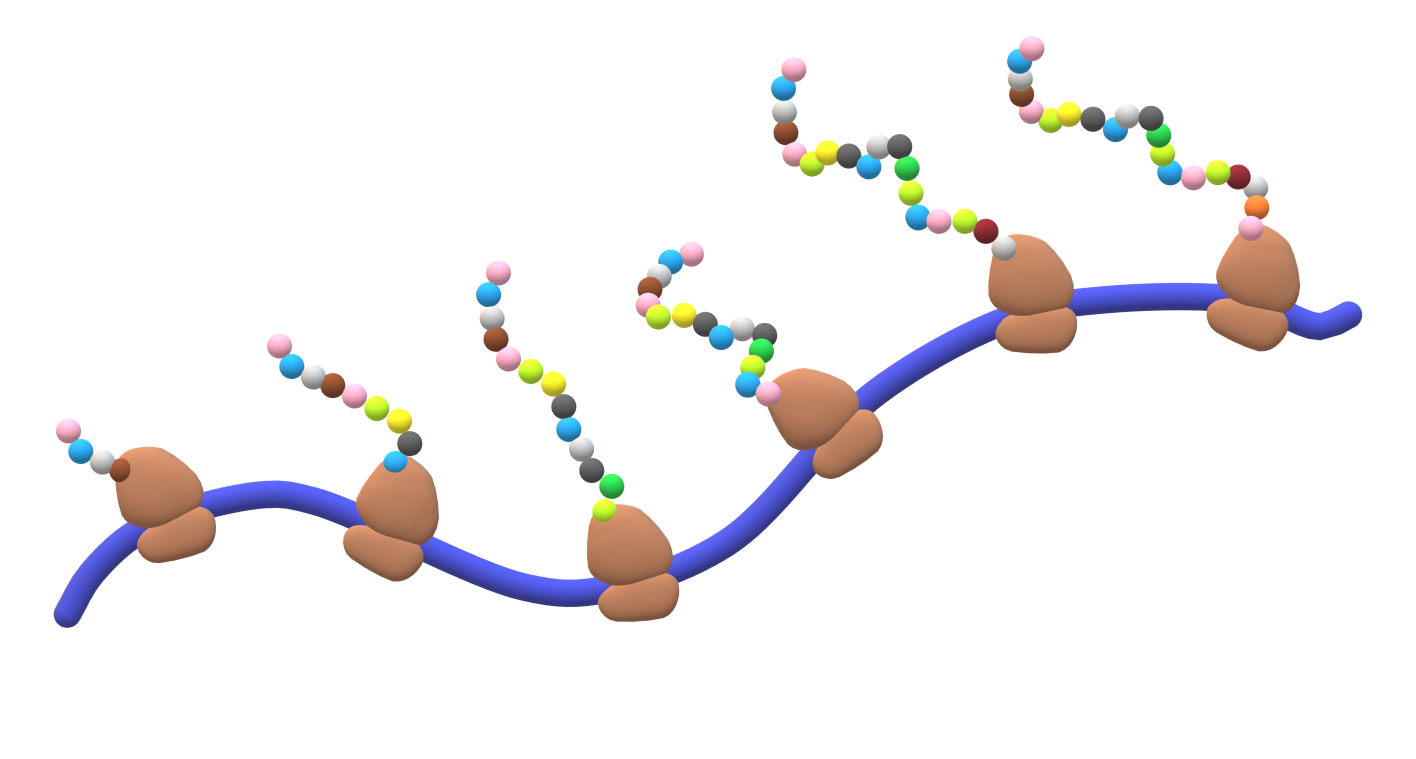
This figure illustrates a polysome: several ribosomes translating the same mRNA concurrently while synthesizing multiple polypeptide chains. It reinforces why polysomes increase protein output—one mRNA can be read repeatedly in parallel, boosting production efficiency. Source
This arrangement:
Amplifies protein production from a single mRNA
Supports rapid responses when a cell needs large amounts of a protein
Ribosomes as evidence of common ancestry
The universality of ribosomes and the conserved nature of:
rRNA
Core ribosomal proteins
The basic translation mechanism (codons, tRNAs, start/stop signals)
all support the concept that modern organisms share an evolutionary origin. Ribosomes are therefore both essential for cellular protein production and an important example of how shared molecular features reflect common ancestry.
FAQ
Many antibiotics bind structural features unique to bacterial ribosomal subunits (70S) and block steps such as tRNA entry or translocation.
Selectivity is not perfect; side effects can occur because mitochondria have ribosomes that share bacterial-like features.
rRNA contributes the core architecture and catalytic activity, including positioning substrates and promoting peptide-bond formation.
Ribosomal proteins mainly stabilise rRNA folding, tune ribosome dynamics, and help with assembly and accuracy.
Accuracy relies on correct codon–anticodon pairing geometry and ribosomal conformational changes that favour correct matches.
Additional proofreading can occur via kinetic checkpoints that reject near-matches before peptide-bond formation.
Differences can arise from translation initiation rates, availability of specific initiation factors, and regulatory proteins or small RNAs that block or enhance ribosome binding.
mRNA localisation and stability also influence how long and where translation occurs.
Targeting information encoded in the polypeptide (often an N-terminal signal sequence) recruits factors that direct the ribosome–mRNA complex to a membrane during early translation.
Without such targeting information, translation typically proceeds on free cytosolic ribosomes.
Practice Questions
State two reasons ribosomes are described as non-membrane structures essential for protein synthesis. (2 marks)
Ribosomes are not surrounded by a membrane / are rRNA–protein complexes rather than membrane-bound organelles (1)
They synthesise polypeptides by translating mRNA / link mRNA codons to amino acids via tRNA (1)
Describe how a ribosome synthesises a polypeptide from an mRNA during translation. (6 marks)
Small subunit binds mRNA and start codon is identified / initiation begins at AUG (1)
Initiator tRNA pairs via anticodon–codon base pairing (1)
Large subunit assembles to form functional ribosome with tRNA-binding sites (1)
During elongation, charged tRNAs enter and match codons in sequence (1)
Peptide bonds form and the ribosome translocates one codon at a time (1)
Stop codon leads to release factor binding and polypeptide release / termination and subunit dissociation (1)

