HL only: A3.2 Classification and cladistics
This entire subtopic is Additional HL content.
Core exam focus: why classification is needed, why traditional taxonomy has limits, how cladistics uses evolutionary relationships, how to read cladograms, and how molecular evidence changed classification.
Why classification is needed
Classification organizes the immense diversity of species into manageable groups.
It makes identification, comparison, communication, prediction and further study easier.
In exams, link classification to shared traits, relationships, and the ability to predict features of organisms in the same group.
Traditional taxonomy vs cladistics
Traditional taxonomy uses ranked groups: kingdom, phylum, class, order, family, genus, species.
Problem: these fixed ranks are arbitrary and do not always match evolutionary divergence.
Groups made using overall similarity can be misleading if similarities are due to convergent evolution rather than common ancestry.
Cladistics is an alternative approach that classifies organisms into clades rather than forcing them into fixed ranks.
A clade contains a common ancestor and all of its descendants.
Best exam phrase: a good classification should correspond to evolutionary relationships.
Advantages of classification based on evolutionary relationships
The ideal classification groups organisms by common ancestry.
Members of the same group should have evolved from a shared ancestor.
If organisms are in the same clade, they are expected to share inherited characteristics.
This allows biologists to predict traits in organisms because related species often share homologous features, genes, or biochemical pathways.
In long-answer questions, link classification + common ancestry + predictive value.
What evidence is used to identify clades?
The most objective evidence comes from base sequences of genes and amino acid sequences of proteins.
Molecular evidence is usually stronger than morphology because it is less likely to be distorted by analogous traits caused by convergent evolution.
Morphological traits can still be used, especially when molecular data are unavailable.
Exam tip: if asked for the best evidence for relatedness, say DNA / RNA base sequences or amino acid sequences.

This diagram is a simple cladogram showing branching relationships between groups. It is useful for practicing how to identify common ancestors, sister taxa, and patterns of divergence. Source
Molecular clock
Sequence differences between groups accumulate gradually over time.
Therefore, the number of differences in DNA base sequences or protein amino acid sequences can be used to estimate when two clades diverged from a common ancestor.
This method is called the molecular clock.
Important limitation: it gives estimates, not exact dates.
Mutation rates can vary with generation time, population size, selective pressure, and other factors.
Exam wording: more sequence differences generally suggest a more distant common ancestor, but only approximately.
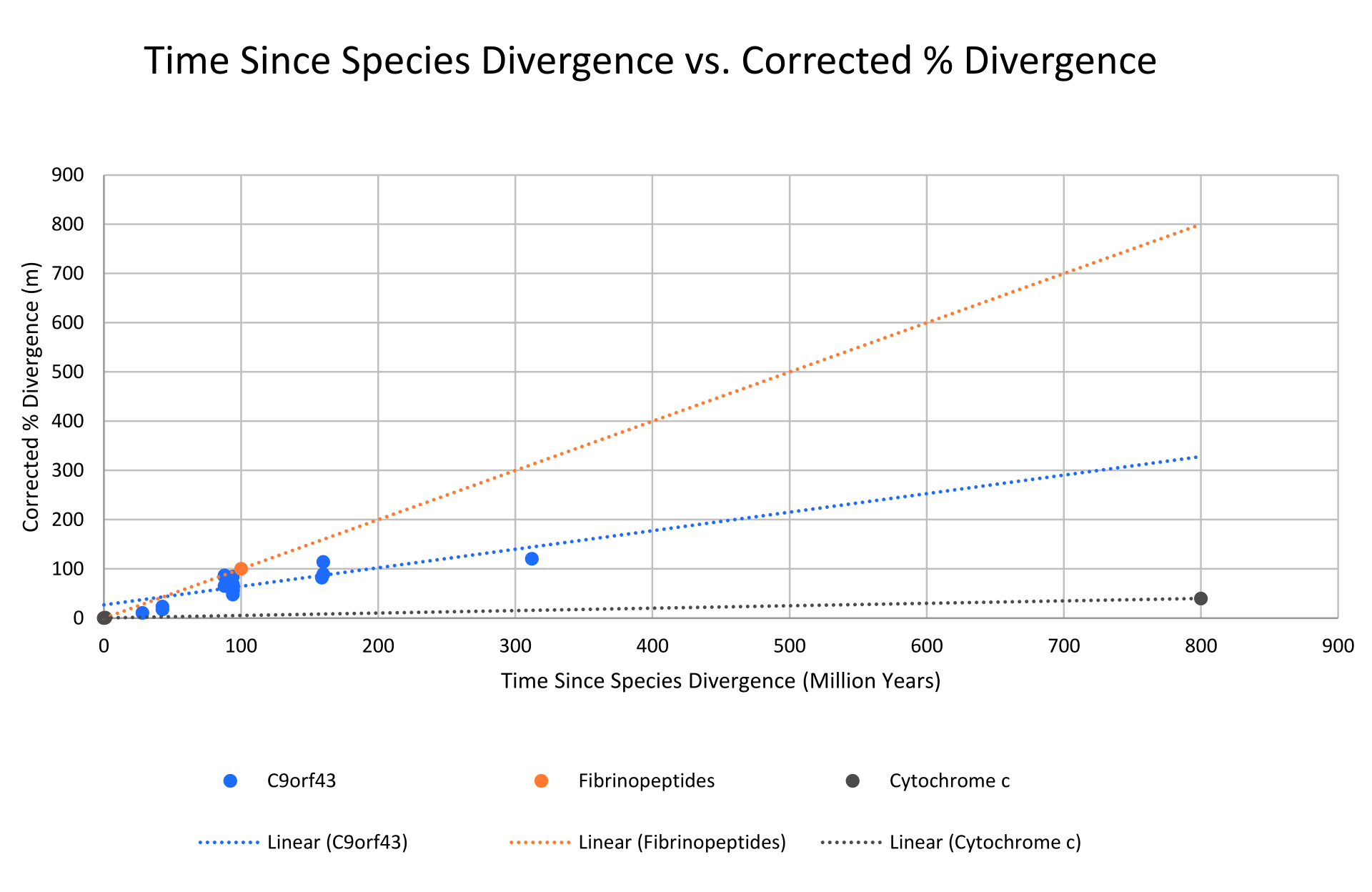
This image shows the idea that sequence changes accumulate over time, allowing approximate estimates of divergence dates. Use it to remember that the molecular clock is comparative and approximate, not exact. Source
Constructing cladograms
Cladograms can be constructed using gene base sequences or amino acid sequences.
The goal is to produce the most probable branching pattern for the taxa being compared.
A common decision rule is parsimony analysis.
Parsimony means choosing the cladogram that requires the smallest number of evolutionary changes.
Different criteria can produce different hypotheses, so cladograms are evidence-based models, not absolute proof.
In data questions, look for the arrangement that explains the observed similarities and differences with the fewest sequence changes.
How to analyse a cladogram
Root = the base of the cladogram; represents the oldest ancestral lineage shown.
Node = a branch point; represents a hypothetical common ancestor.
Terminal branch = the branch ending in a named taxon; represents the descendant group being compared.
Taxa that share a more recent node are more closely related.
A clade consists of a node and all descendants from that node.
Cladograms show patterns of relatedness, not “progress” or “advancement”.
The order of taxa across the tips may rotate without changing the relationships.
To find the closest relatives, trace backwards until two taxa meet at their most recent common ancestor.
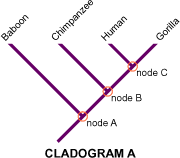
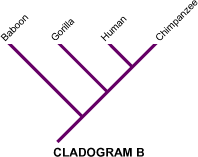
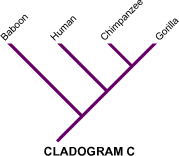
These diagrams are excellent for learning how to identify nodes, sister taxa, outgroups, and alternative hypotheses of relationship. They also reinforce that a cladogram is a testable hypothesis built from evidence. Source
Using cladistics to test existing classifications
Cladistics can test whether traditional groups really reflect evolutionary relationships.
Sometimes organisms were grouped together because of morphological similarity, but later molecular evidence showed they were not close relatives.
This often happens when similarity is due to convergent evolution.
So cladistics can falsify older classifications and lead to reclassification.
You do not need to memorize the details of the figwort family (Scrophulariaceae) case study.
What you do need: understand the principle that new molecular evidence can change classification.
Three-domain classification
Evidence from rRNA base sequences led to classification of all organisms into three domains:
Bacteria
Archaea
Eukarya
This added a level above kingdoms and was a major shift in biological classification.
Key point: rRNA sequences were important because they revealed deep evolutionary differences not obvious from appearance alone.
In exam answers, connect molecular evidence to the revolutionary reclassification of life.
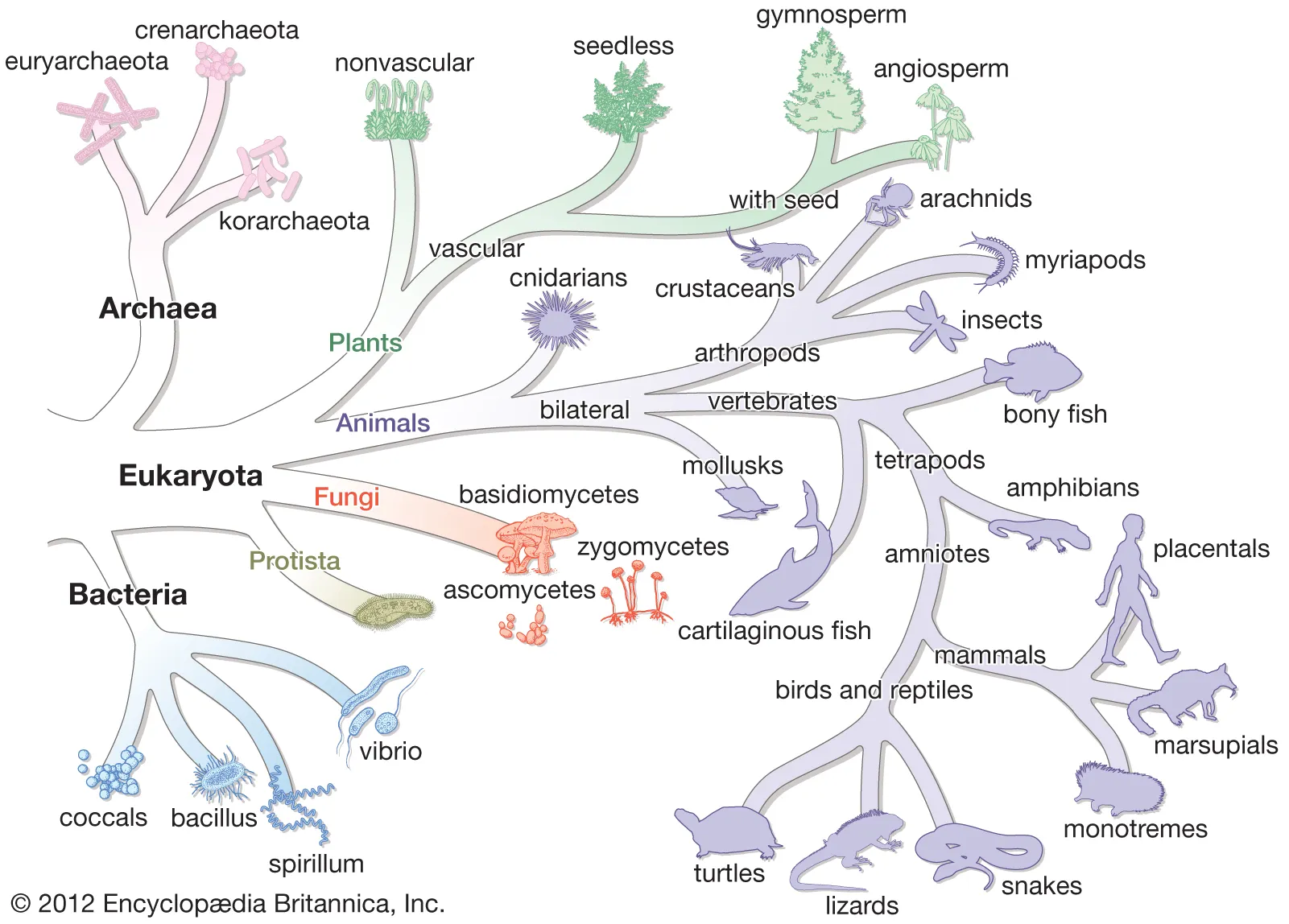
This figure shows the separation of life into Bacteria, Archaea, and Eukarya based on molecular evidence. It supports the idea that rRNA sequence data transformed biological classification. Source
Nature of science links that matter in exams
Fixed taxonomic ranks are arbitrary: they do not perfectly reflect the continuous nature of evolutionary divergence.
Cladistics represents a paradigm shift because classification moved from ranked similarity-based groupings toward evolutionary, evidence-based clades.
Different criteria can produce different hypotheses: for example, different analyses may give different cladograms.
Parsimony is one criterion for choosing the most probable cladogram.
Scientific classifications can be revised or falsified when new evidence appears.
Similar morphology does not always mean close relationship because of convergent evolution.
Common exam traps
Do not say organisms are related because they “look similar” unless the question specifically supports that with evidence.
Do not confuse a clade with any arbitrary group of similar organisms.
Do not treat a node as a real observed species; it represents a hypothetical common ancestor.
Do not assume fewer visible differences always means closer relationship; molecular data may show otherwise.
Do not claim the molecular clock gives an exact divergence date.
Do not read a cladogram from left to right as a ladder of “higher” and “lower” organisms.
Checklist: can you do this?
Explain why cladistics is often better than traditional ranked taxonomy for showing evolutionary relationships.
Identify root, node, terminal branch, clade, common ancestor, and sister taxa on a cladogram.
Deduce which taxa are most closely related and which shared a more recent common ancestor.
Interpret how DNA / protein sequence differences and the molecular clock are used to infer divergence.
Apply parsimony to choose the most likely cladogram from simple exam data.
Ultra-compact exam summary
Classification is needed to organize biodiversity.
Traditional ranked taxa are useful but can be arbitrary and may not match evolutionary history.
Cladistics groups organisms into clades based on common ancestry.
Strongest evidence for clades = gene base sequences and amino acid sequences.
Molecular clock uses accumulated sequence differences to estimate divergence time.
Parsimony chooses the cladogram with the fewest changes.
In a cladogram: node = hypothetical common ancestor, root = oldest lineage shown, terminal branch = descendant taxon.
rRNA evidence led to the three domains: Bacteria, Archaea, Eukarya.

Shubhi is a seasoned educational specialist with a sharp focus on IB, A-level, GCSE, AP, and MCAT sciences. With 6+ years of expertise, she excels in advanced curriculum guidance and creating precise educational resources, ensuring expert instruction and deep student comprehension of complex science concepts.
Shubhi is a seasoned educational specialist with a sharp focus on IB, A-level, GCSE, AP, and MCAT sciences. With 6+ years of expertise, she excels in advanced curriculum guidance and creating precise educational resources, ensuring expert instruction and deep student comprehension of complex science concepts.